CLEARED FOR TAKEOFF
BY JENNIFER BARBER • PHOTO COURTESY GLOBAL INSTITUTE FOR FOOD SECURITY
Crop breeders test thousands of plant lines every year in small, individual test plots. Assessing these plants involves both quantitative and qualitative analysis, but new software aims to substantially refine the process. PlotVision is a new software service that collects data using unmanned aerial imagery (UAI) captured by drones. The data may help researchers predict harvest yield and assess disease resistance, accelerate the plant breeding process and the production of new crop varieties.
The PlotVision software was developed through the Plant Phenotyping and Imaging Research Centre (P2IRC), which is run by the Global Institute for Food Security (GIFS) on behalf of the University of Saskatchewan (USask). The software translates the work of the P2IRC program into a practical format that can be used by plant breeders.
The P2IRC program benefits from the technical expertise and infrastructure available at USask, and this includes its Crop Development Centre. Engineers work with computer scientists, who work with researchers in the university’s College of Agriculture and Bioresources. This infrastructure facilitates the development of new varieties through to commercial availability.
The PlotVision project was created in 2019 to examine the physical characteristics of plants to see how they perform under certain conditions. It has been a collaborative effort between plant breeders and computer scientists.
“The program is a community effort,” said Chris Barker, director of research and business development for P2IRC and GIFS. “While it is managed by the GIFS, there is great work being done throughout the university, as well as with Agriculture and Agri-Food Canada (AAFC) and with our other partners. It truly has been a multidisciplinary effort.”
“The ultimate goal of PlotVision is to do more on the computer and less in the field to support the plant breeder with making better decisions faster,” said Barker. “It’s not supposed to replace the role of the plant breeder, but is rather intended to combine with traditional on-ground data collection. It can provide additional quantitative data to aid breeders’ decision-making, which will then allow them to focus on other areas of seed development.”
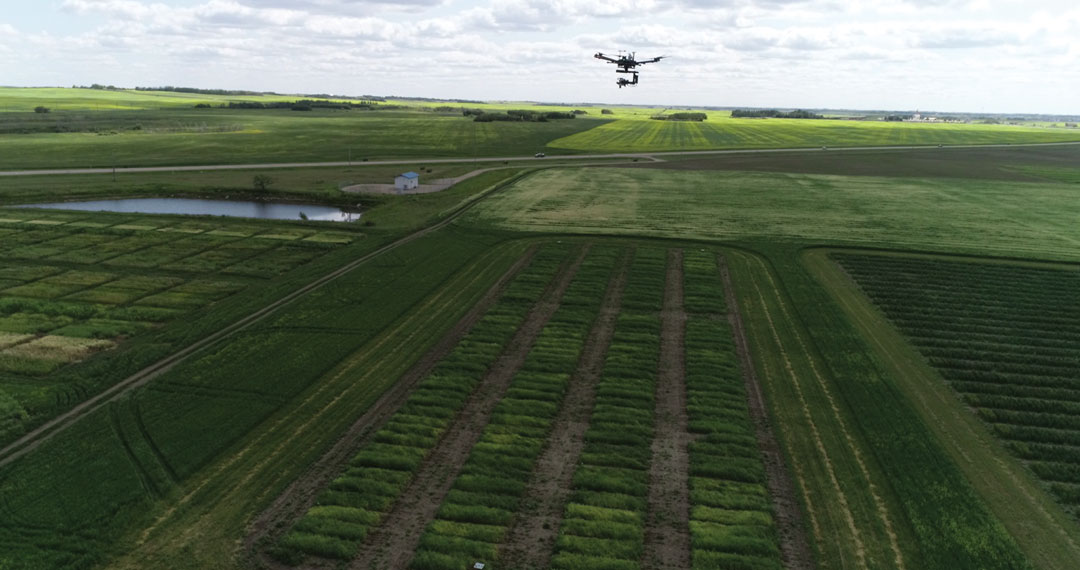
The software was developed by P2IRC research associate William van der Kamp, under the leadership of USask computer scientist Ian Stavness. A drone takes images over a test plot and the software uses artificial intelligence (AI) to convert the image into useful information for breeders. PlotVision has been used to analyze leaf and flower colour to gauge maturity, and uses the 3D shape to assess crop architecture and canopy volume. Understanding these characteristics assists in predicting crop outcomes. Researchers can use this information to identify promising crop lines as well as best management practices.
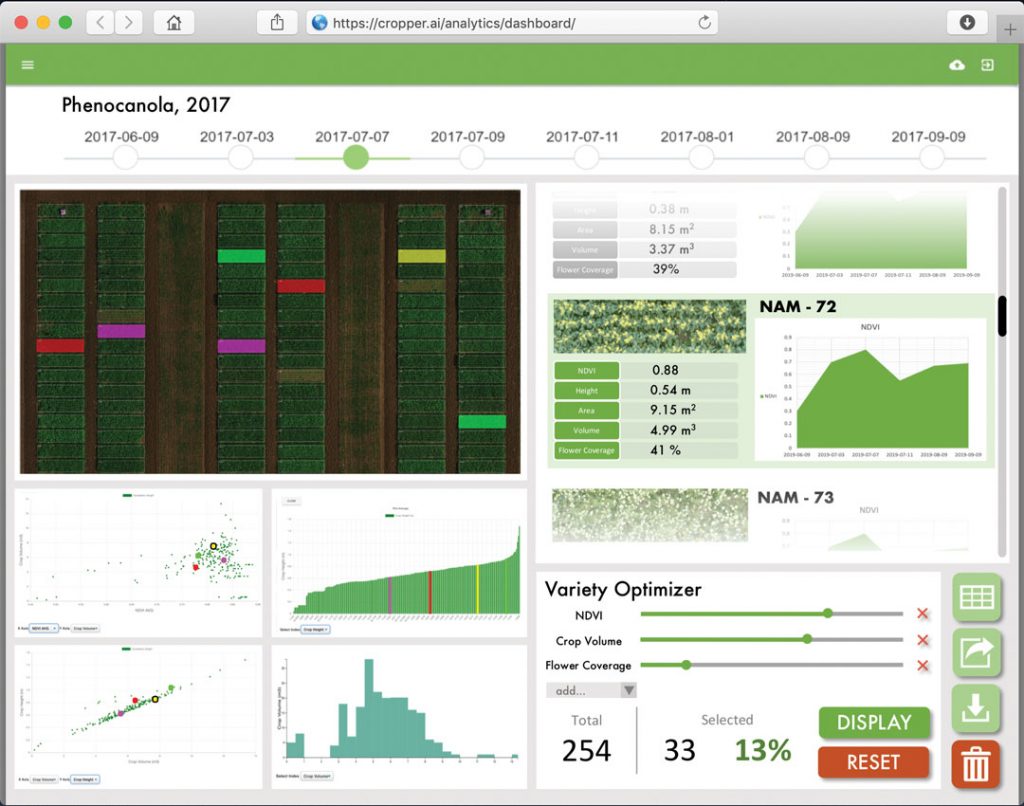
Among the AI tools used by PlotVision are semantic segmentation neural networks. These digital systems are able to identify which pixels in an image represent various objects, such as soil and plants. Establishing which pixels in an image represent soil, it can then be eliminated from the analysis. PlotVision also uses other types of AI technology.
With enough training, these software components learn to count individual plants as well as identify leaves and dead plant matter. Importantly, the system identifies material at a much smaller scale than satellite imagery and produces very refined results.
“Images represent data, and PlotVision serves as a tool to translate large collections of plant data in order to get very specific information for breeders,” said van der Kamp. “The images on their own are of little value, but this tool helps us to build more and more analytics around them for breeders to use in their test plots. For example, breeders may count 60 to 75 seedlings from an image of a small, early stage canola plant. But using an algorithm through PlotVision, they can build on those numbers and automatically perform many more seedling counts to get much more statistically relevant information.”
Currently, very large numbers of field plots must be manually assessed over an approximately seven-year cycle to develop each new commercial variety. PlotVision will help make this process more efficient, and will improve the quality of the assessments. The analysis is automated and data-driven, eliminating the potential for human error, while reducing costs and turnaround time for analysis.
The software was initially used to test canola, lentils and wheat, as these are key rotational crops in Saskatchewan. Van der Kamp said once the AI and the analytics component of the system have been fully tested the system should also be applicable to additional crops.
PlotVision is a natural fit with the P2IRC program’s plant breeding focus, as the tool in turn focuses on the individual plant lines in breeders’ structured field designs, explained van der Kamp. “Working through P2IRC’s plant breeding program to develop this technology made practical sense because testing areas in the program incorporate many plant lines in replicated plots with known genotypes, all in a relatively small area, said van der Kamp. “The images and resulting data we derive through PlotVision provide very robust information to help breeders make important decisions.”
For the 2020 growing season, project participants planned to work with local breeding programs at USask and with AAFC to further expand the capabilities of the software, including what it is able to identify and how its data can be utilized. “Though the size of the field program was impacted by the COVID-19 pandemic, PlotVision’s many features continue to be validated, providing valuable information to our breeding partners and great benefit to farmers and the agriculture industry,” said Barker.
In 2021, PlotVision will be made available for use beyond P2IRC, and will eventually be offered on a fee-for-service basis through the GIFS-managed Omics and Precision Agriculture Laboratory.








Comments